How Does Genomic Medicine Become a Reality for Children’s Health?
How Does Genomic Medicine Become a Reality for Children’s Health? https://pediatricsnationwide.org/wp-content/uploads/2020/11/Achieving-Clarity-Header-1024x575.jpg 1024 575 Jan Arthur Jan Arthur https://secure.gravatar.com/avatar/6acc85c9c2e25aeffffc346df0fa47b9d86773e3ce0d64a7a7f65c936791f7b1?s=96&d=mm&r=g
Perhaps one of the most important initiatives to advance our understanding of pediatric disease is genomic analysis.
Genomics encompasses all aspects of understanding the human genetic code, especially how genetic information contained in every human cell can be interpreted to prevent disease and customize treatments for nearly every type of illness.
Genomic analysis allows us to search for the cause of a child’s illness by looking for variations within their genetic code. To find those variations, the child’s genetic code is compared to a reference code generated by analyzing other individuals without the disease in question. It’s within those variations that researchers can answer key questions regarding health and disease for individual patients to entire populations.
However, that step isn’t simple.
Making Genomic Medicine Viable
Imagine looking for one typo in a 100-volume book collection. That’s genomic analysis — to search for that one genetic variation that may cause disease, to discover an effective gene-based therapy or to explain genetic trends in entire populations. The answer to a patient’s search could be as hidden and difficult to find as that one misspelled word in a library of books.
More than a decade ago in a watershed moment in our scientific history, the Human Genome Project was completed, sequencing an entire human genome. It took 15 years and about $3 billion to accomplish. Today, sequencing that same human genetic blueprint takes about one day and costs around $2000. And it’s considered a somewhat ho-hum feat.
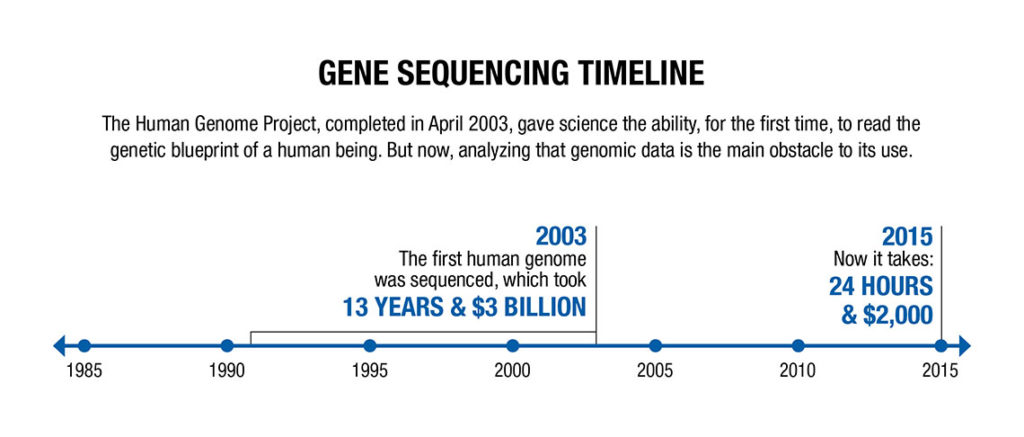
However, this unassuming feat still generates billions of data points that need to be analyzed before any misspellings can be found. Analyzing that data in the attempt to make it useful has proved to be daunting. Recently, best-in-class technologies have decreased the length of time to three days for analysis.
What may be surprising to some, a research lab at a Midwestern children’s hospital has developed an even faster and more accurate analytic solution – one that can truly make genomic medicine useful in clinical and research settings.
An algorithm named Churchill was created by Peter White, PhD, director of the Biomedical Genomics Core at Nationwide Children’s, and his team. With this computational pipeline, analyzing a single human genome takes about 90 minutes and costs less than $1000.
With that efficiency, analysis of larger populations also becomes a tangible goal.
In a paper published in the January 2015 edition of Genome Biology, Churchill demonstrated its capabilities for population-scale analysis by re-analyzing the genomic data from the 1,000 Genomes Project. Through an award from the Amazon Web Services (AWS) in Education program, Dr. White and his team successfully analyzed phase 1 of the raw data generated by the 1000 Genomes Project in a fraction of the original time.
The 1000 Genomes Project was an international collaboration to produce an extensive public catalog of human genetic variation, representing multiple populations from around the globe. It required 18 months, more than 400 scientists and $10 million to complete the analysis.
Using cloud-computing resources from AWS, Churchill completed analysis of the 1,088 whole genome samples in seven days with five scientists at a cost of about $70,000. In addition, Churchill identified millions of new genetic variants not found through the 1000 Genomes Project.
Churchill’s output was validated using National Institute of Standards and Technology (NIST) benchmarks. In comparison with other computational pipelines, Churchill was shown to have the highest sensitivity at 99.7 percent; highest accuracy at 99.99 percent and the highest overall diagnostic effectiveness at 99.66 percent. It’s also 100 percent reproducible. The technology has been licensed to GenomeNext LLC for population scale analyses.
With all this computational power, researchers can now turn toward the question of “what next?”.
Using Genomics to Solve Diseases
In pediatrics, many devastating diseases are rare and often have unknown origins. With genomic analysis becoming more clinically practical, clinicians may be able to more efficiently identify the genetic variations causing a disease. In many cases, additional research is then needed to identify ways to actually treat that disease.
In 2015, Boston Children’s Hospital presented a challenge – open to any organization – to unravel answers for five patients faced with debilitating diseases with no known cause or cure. Twenty-six institutions from around the world participated in the CLARITY Undiagnosed challenge. Based on the capabilities of Nationwide Children’s Churchill technology, the diverse team of clinicians and scientists and their comprehensive approach, the hospital won the challenge.
For more familiar diseases such as pediatric cancers and neuromuscular disorders, genomic research and its associated gene therapies are inching closer to the potential for cures.
Identifying Genes to Generate Cures
The ability to implicate pathogenic genes is beginning to demonstrate promise for gene-based therapies. For neuromuscular disorders, Nationwide Children’s is conducting more gene therapy clinical trials than any other institution in the world.
Spinal muscular atrophy type 1, a degenerative neuromuscular disease, is the most common genetic cause of death for infants. Virtually all children affected with SMA type 1 die by 2 years of age.
In 2014, Jerry Mendell, MD, director of the Center for Gene Therapy at Nationwide Children’s, began a phase 1 gene transfer clinical trial for SMA type 1. In this trial, a large number of modified viruses containing the missing survival of motor neuron 1 (SMN1) gene are administered to the patient. By administering the gene early in the disease process, the team aims to effectively halt the progression of the disease.
Sanfilippo syndrome type A is another rare condition where gene therapy may be the answer. The syndrome is a rare and life-threatening genetic disease that leads children to progressively lose the ability to speak, walk and eat before their eventual deaths. Children appear healthy at birth and usually do not show symptoms until after their first year of life. Currently, there is no cure for Sanfilippo syndrome and treatments mainly involve supportive care.
In February, Abeona Therapeutics, a company focusing on therapies for life-threatening rare diseases, announced FDA approval of a new clinical trial for Sanfilippo syndrome type A. The trial will use a gene therapy approach, also known as ABO-102, developed by researchers at Nationwide Children’s.
Expanding Capabilities for the Future
Renowned scientists Richard Wilson, PhD, and Elaine Mardis, PhD, currently lead the McDonnell Genome Institute at Washington University, one of only four genomics centers funded by the National Human Genome Research Institute.
They also believe in the potential of genomic medicine.
Their work has already led to a number of key findings, including changing the course of therapy for a deadly form of leukemia, uncovering a drug target in a form of eye cancer and performing the first clinical trial of personalized vaccines for melanoma patients.
In 2008, their team became the first to use DNA sequencing technology to compare the tumor DNA of a cancer patient with that patient’s normal tissue DNA. It demonstrated that genetic differences between tumor and normal gene sequences could identify mutations driving cancer growth. This foundational work resulted in an international effort to decode cancer genomes to improve treatments and outcomes.
Beginning in the fall of 2016, Dr. Wilson and Dr. Mardis will move to Nationwide Children’s to continue their work, specifically aimed at pediatric diseases.
“We’ve been fortunate to play a role in the rapid advance of modern genomics, especially the use of next-generation sequencing (NGS) to enable better understanding and diagnosis of human disease,” said Dr. Mardis. “This has been particularly inspiring for us with regard to the impact on a variety of pediatric conditions, ranging from developmental delay to metabolic deficiencies to cancer. We feel strongly that the application of NGS to diagnoses can add precision to therapeutic decision-making and other diagnostic assays.”
In a recent interview published GenomeWeb, the team explained that their main goal will be to combine research and clinical efforts in order to have the smoothest transition from what is learned in the research setting into real-time clinical use.
In the long term, they note, genomics needs to demonstrate its critical role in precision diagnostics and clinical benefit so that insurers reimburse its use and health practitioners recognize its importance to precision health.
Nationwide Children’s is investing in the promise of genomics to have an unprecedented role in the diagnosis, treatment and prevention of pediatric disease, certain that its role will make a significant difference in the future of child health.





