A Brave New World of Data
A Brave New World of Data https://pediatricsnationwide.org/wp-content/themes/corpus/images/empty/thumbnail.jpg 150 150 Katie Brind'Amour, PhD, MS, CHES Katie Brind'Amour, PhD, MS, CHES https://pediatricsnationwide.org/wp-content/uploads/2021/03/Katie-B-portrait.gifAccording to William C. Ray, PhD, of the Battelle Center for Mathematical Medicine in The Research Institute at Nationwide Children’s Hospital, novel techniques prototyped for understanding protein data could soon enable the visualization of large sets of demographic data, displaying contingency tables and group relationships through visual representations of positive and negative correlation, covariance and mutual exclusivity.Massive, multi-dimensional contingency tables are a ubiquitous feature of biological data. From precision medicine to structural biophysics, understanding complex networks of dependencies in contingency data is critical to drawing knowledge from and making informed decisions about the data.

Protein data is often shown in this classic sequence alignment, with strings of letters for each amino acid in the order a cell assembles them.
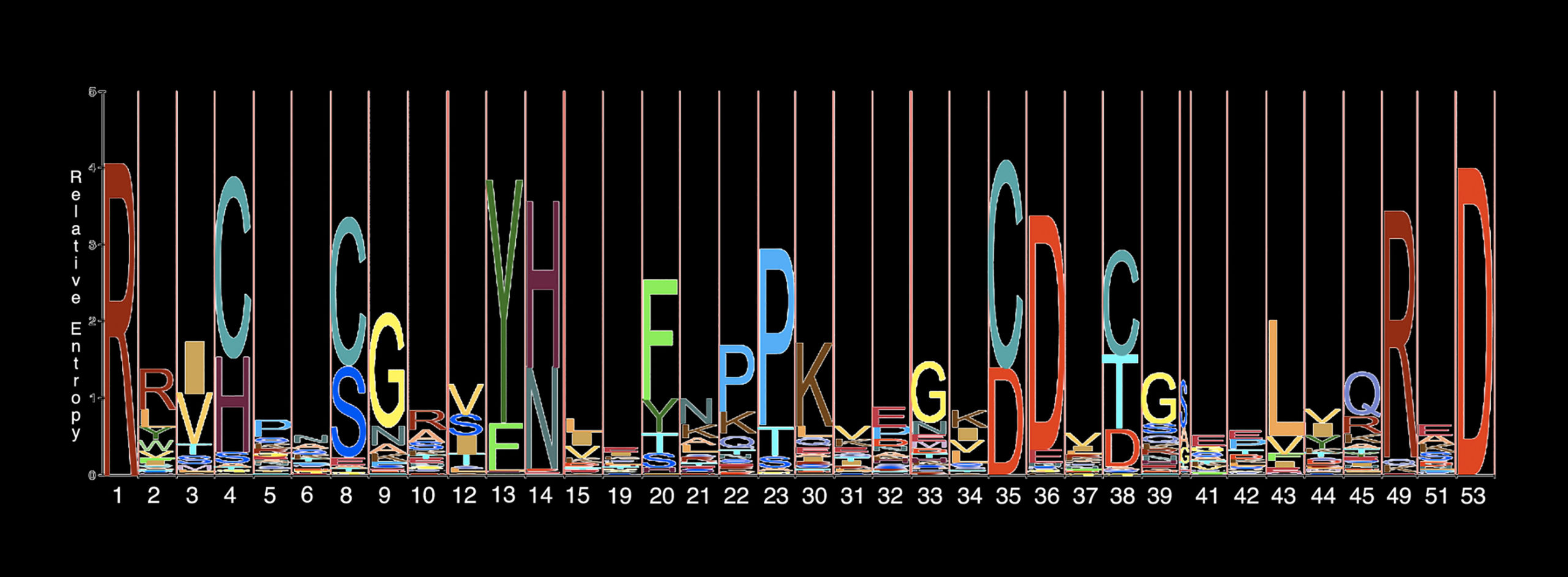
Sequence logos depict a protein’s amino acid sequences but lack information about its structure.
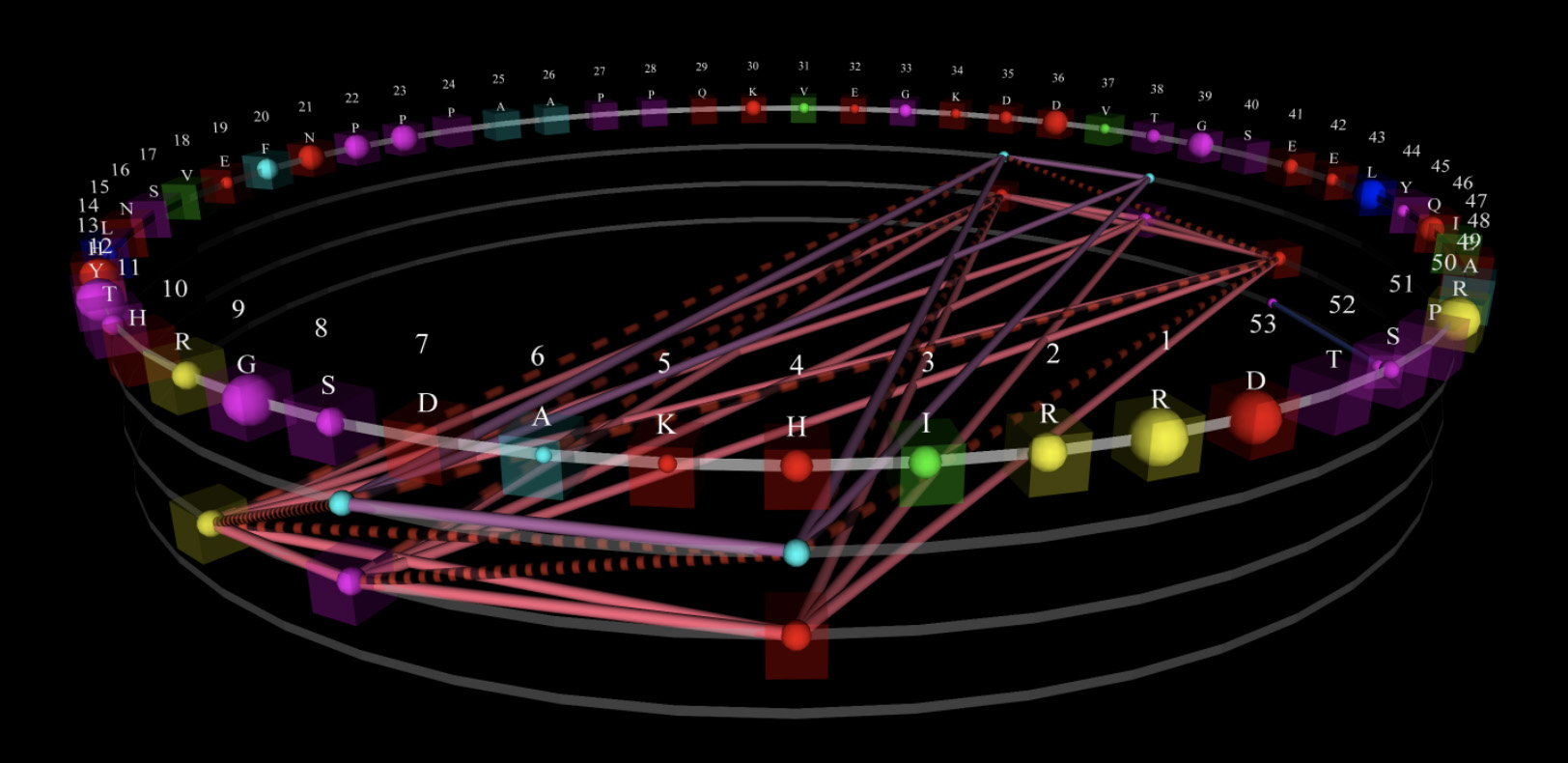
The many appearances of the Adenylate Kinase (ADK) protein lid domain and the inability of character sequences to show functional dependencies between the amino acid subunits highlight the need for data representations that are more sophisticated. But existing methods of creating precise molecular models are laborious, expensive and rarely successfully executed.
This visualization of ADK reveals that, although certain amino acids are far apart in the protein sequence, they are nearby in the physical structure of the model. The data representation also identifies previously unknown amino acid interactions. StickWRLD models offer insight into structure and function requirements for proteins and hold similar potential for other fields with complex, interacting data.
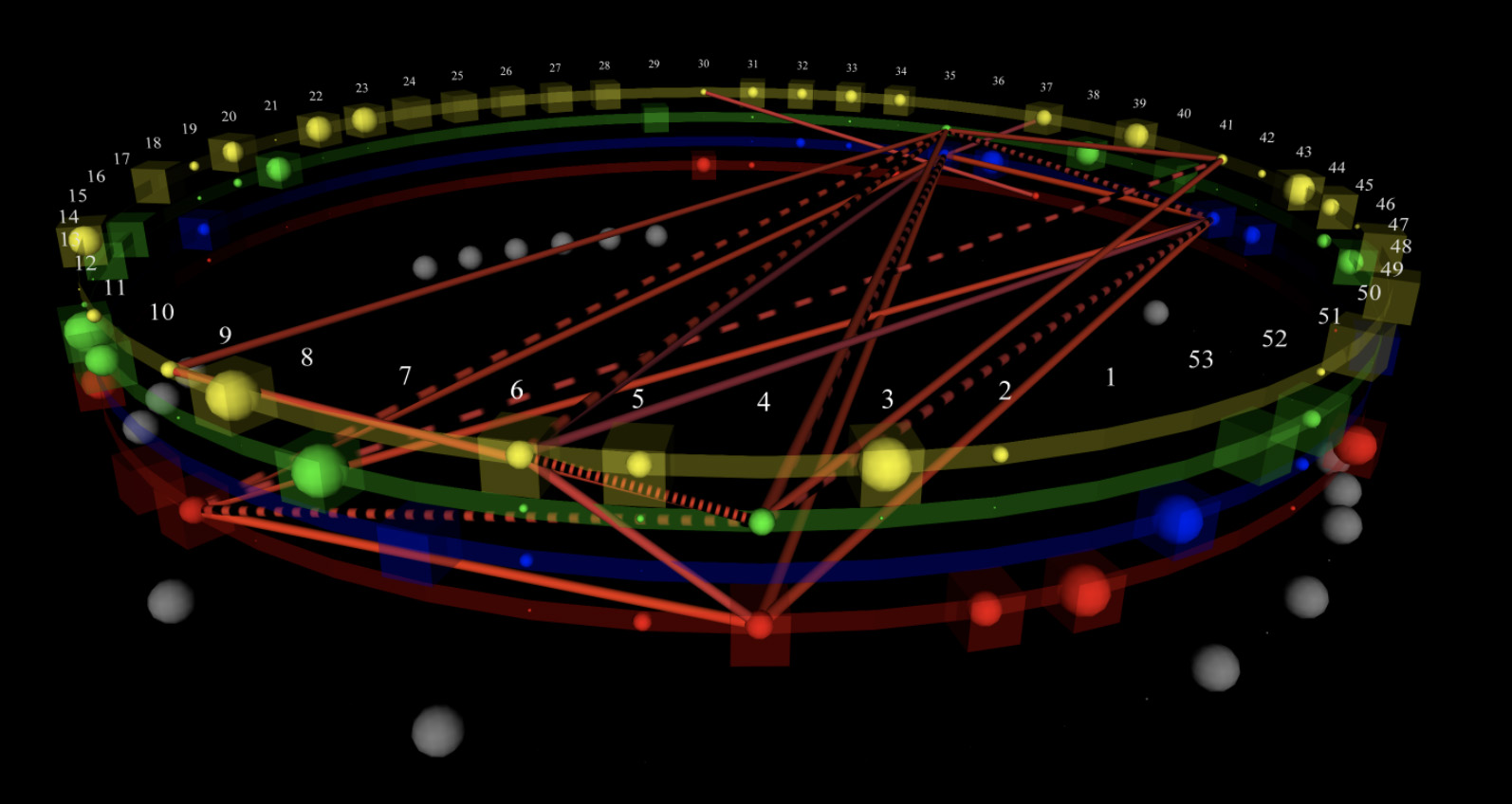
Alternative representations enable amino acid grouping by physical categories to see whether relationships depend on properties such as charge or size.
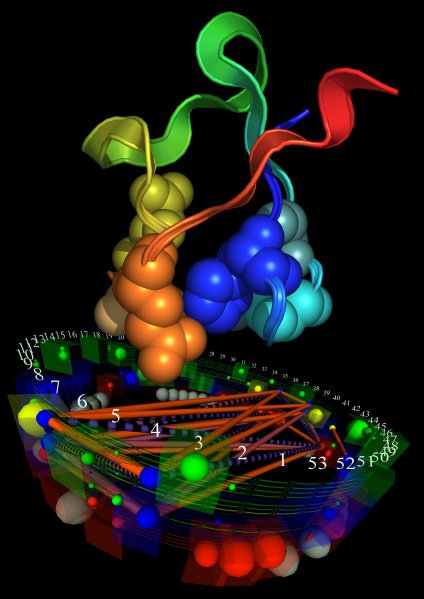
Visually representing ADK data with this program allows researchers to understand how multiple species can share an identical molecular configuration that has different ways of maintaining its structure and function.
The story appeared in the Spring/Summer 2014 print issue. Download the full issue.
About the author
Katherine (Katie) Brind’Amour is a freelance medical and health science writer based in Pennsylvania. She has written about nearly every therapeutic area for patients, doctors and the general public. Dr. Brind’Amour specializes in health literacy and patient education. She completed her BS and MS degrees in Biology at Arizona State University and her PhD in Health Services Management and Policy at The Ohio State University. She is a Certified Health Education Specialist and is interested in health promotion via health programs and the communication of medical information.
- Katie Brind'Amour, PhD, MS, CHEShttps://pediatricsnationwide.org/author/katie-brindamour-phd-ms-ches/April 27, 2014
- Katie Brind'Amour, PhD, MS, CHEShttps://pediatricsnationwide.org/author/katie-brindamour-phd-ms-ches/April 27, 2014
- Katie Brind'Amour, PhD, MS, CHEShttps://pediatricsnationwide.org/author/katie-brindamour-phd-ms-ches/April 27, 2014
- Katie Brind'Amour, PhD, MS, CHEShttps://pediatricsnationwide.org/author/katie-brindamour-phd-ms-ches/April 28, 2014




